Chemical group transformations¶
SILCS-MC ligand optimization and SSFEP both entail computations on derivatives of a parent ligand. Additionally, the user may wish to perform SILCS docking using a series of structures chemically related to a reference compound (i.e. parent ligand). Therefore, a task common to these use cases is that of performing chemical group transformations on a parent ligand to create a series of chemically-related compounds.
The SilcsBio GUI provides an intuitive way to handle this task directly in the context of SILCS-MC ligand optimization (see Running SILCS-MC ligand optimization from the SilcsBio GUI) and SSFEP (see SSFEP simulation from the GUI). You can also use the SilcsBio GUI to perform chemical transformations independently of any other task by choosing Modify ligand for SILCS-MC/SSFEP from the Home page. Output Mol2 files will be saved to a directory of your choosing on the machine where you are running the SilcsBio GUI.
It is also possible to perform chemical group transformations using the command line interface, as described below.
You will need to create a text file modifications.txt with instructions
listing the desired modifications to your parent ligand.
Three types of modifications can be performed, as listed in the table.
Command |
Modification |
|---|---|
JOIN |
Join a fragment (mol2) to the parent at a defined site |
REPL |
Replace an atom at a defined site on the parent with a fragment (mol2) |
MUDE |
Change an atom at a defined site on the parent to another atom-type |
An example ligand modification input file is povided with your SilcsBio
server software, ${SILCSBIODIR}/examples/ssfep/modification.txt.
For details on how to modify and process this file to create Mol2 files containing modifications to your parent ligand, run the following command:
${SILCSBIODIR}/programs/chemmod -h
Your SilcsBio server software comes with a large number of fragments in Mol2 format that can be added/joined to your parent ligand. These fragments are found in:
${SILCSBIODIR}/data/groups
Modifications JOIN example¶
A JOIN operation can be performed between the parent and a ch4.mol2
fragment by adding the following line to modifications.txt:
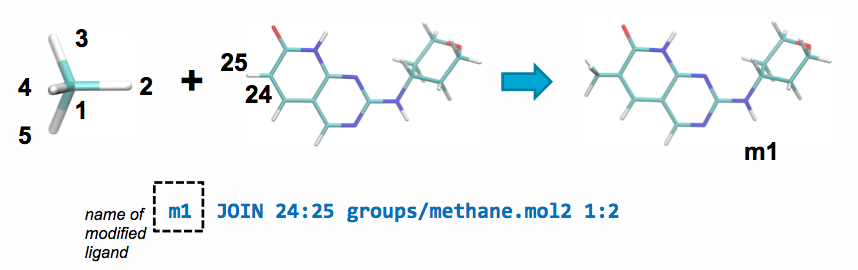
This line will join atom 24 in the parent ligand with
atom 1 in methane.mol2 and delete atoms 25 and 2 in the parent ligand and
the joined fragment, respectively.
Modifications REPL example¶
A REPL operation can be performed between the parent and a ch4.mol2
fragment by adding the following line to modifications.txt:
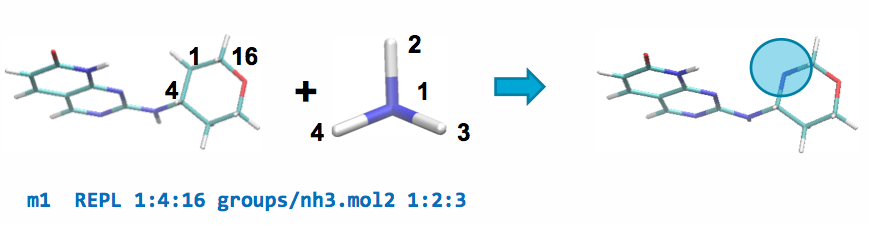
This line will replace atom 1 of the parent ligand with atom 1 of the fragment by aligning atoms 2 & 3 of the fragment with atoms 4 & 16 of the parent, respectively, and replacing the carbon in the ring with a nitrogen atom.
Modifications MUDE example¶
The same transformation in the previous section can also be achieved using a MUDE operation:
m2 MUDE MU 1:7 DE 21
This line will mutate atom 1 (atom index number) in the parent with nitrogen (atom index number 7) along with deleting the hydrogen (atom index number 21) attached to the parent carbon.
Ligand decoration¶
To evaluate multiple modifications to a single site on the parent ligand, use the following syntax:
m1 JOIN 24:25 <filename>
Replace <filename> with the name of a text file containing each of the
multiple modifications, with one modifications per line. Examples of such
text files are list.txt and small_join_list.txt located in the
${SILCSBIODIR}/data/ folder, and you can use these example files as
the basis for your own custom file that includes modifications using Mol2
files from ${SILCSBIODIR}/data/groups or your own custom Mol2 files.
Be careful to pay attention to chemistry; if a modification in your text
file is not suitable for a site, you can comment it out using ! at
the beginning of that line. Based on the above JOIN line, Mol2 output
files from running ${SILCSBIODIR}/programs/chemmod will all begin
with the prefix m1_.